I received my Ph.D. in 2023 from the University of Chinese Academy of Sciences and am currently conducting postdoctoral research at the University of Florida. My research focuses on computational drug discovery and molecular design, combining computational chemistry, computational biology, and artificial intelligence to accelerate protein engineering and small-molecule drug discovery.
My current research interests include:
- generative AI for de novo protein design;
- virtual screening and molecular design based on CADD and AIDD;
- molecular mechanism studies of enzymes and GPCRs using molecular simulations.
🔥 News
- 2025/12/26: Our research article Sacrificing Knight for Pawn-Queen Promotion: Sonosensitive Nano-Xanthiums Orchestrate NO to Retain H2O2 for Antibiofilm Therapy was published in Advanced Materials.
- 2025/05/28: Elected as a member of Sigma Xi, The Scientific Research Honor Society.
- 2025/03/26: Served as a reviewer for ACS journals and contributed to international academic publishing.
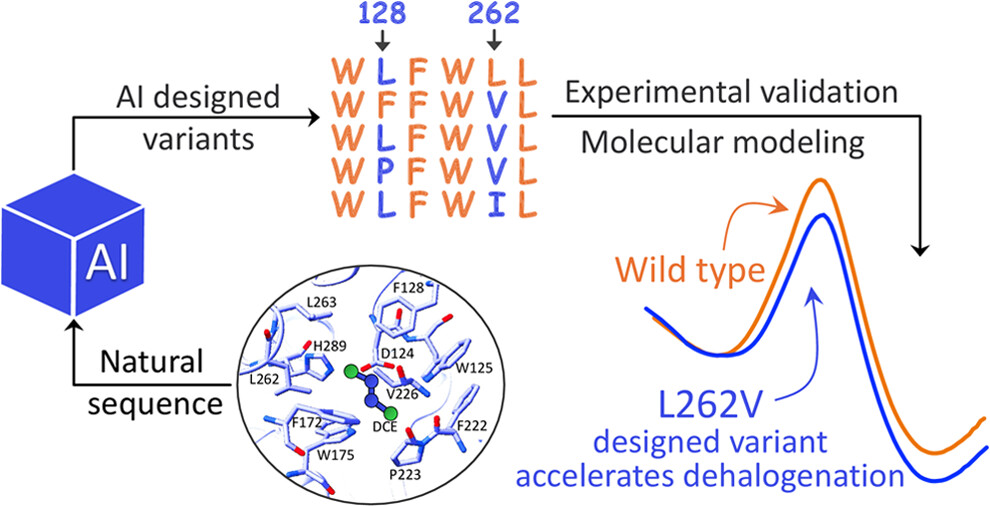
- 2025/01/10: Our research article Biochemical and Computational Characterization of Haloalkane Dehalogenase Variants Designed by Generative AI: Accelerating the SN2 Step was published in Journal of the American Chemical Society.
- 2024/04/10: Joined the Chinese Chemical Society and continued participating in academic exchange.
- 2024/03/03: Our review article Will the hype of automated drug discovery finally be realized? was published in Expert Opinion on Drug Discovery.
- 2024/02/15: Our review article Computational drug development for membrane protein targets was published in Nature Biotechnology.
📝 Publications
Biochemical and Computational Characterization of Haloalkane Dehalogenase Variants Designed by Generative AI: Accelerating the SN2 Step
Natalia Gelfand1, Vojtech Orel1, Wenqiang Cui1, Jiri Damborsky, Chenglong Li, Zbynek Prokop, Wen Jun Xie, Arieh Warshel
- A representative study on using generative AI to support enzyme and protein design with combined biochemical and computational validation.
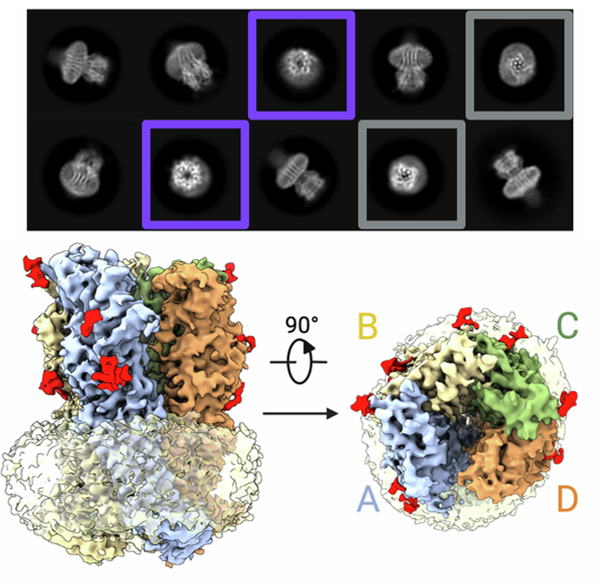
Structure of tetrameric forms of the serotonin-gated 5-HT3A receptor ion channel
Bianca Introini1, Wenqiang Cui1, Xiaofeng Chu1, Yingyi Zhang, Ana Catarina Alves, Luise Eckhardt-Strelau, Sabrina Golusik, Menno Tol, Horst Vogel, Shuguang Yuan, Mikhail Kudryashev
- A structural study revealing tetrameric forms of the serotonin-gated 5-HT3A receptor ion channel and their implications for receptor assembly.
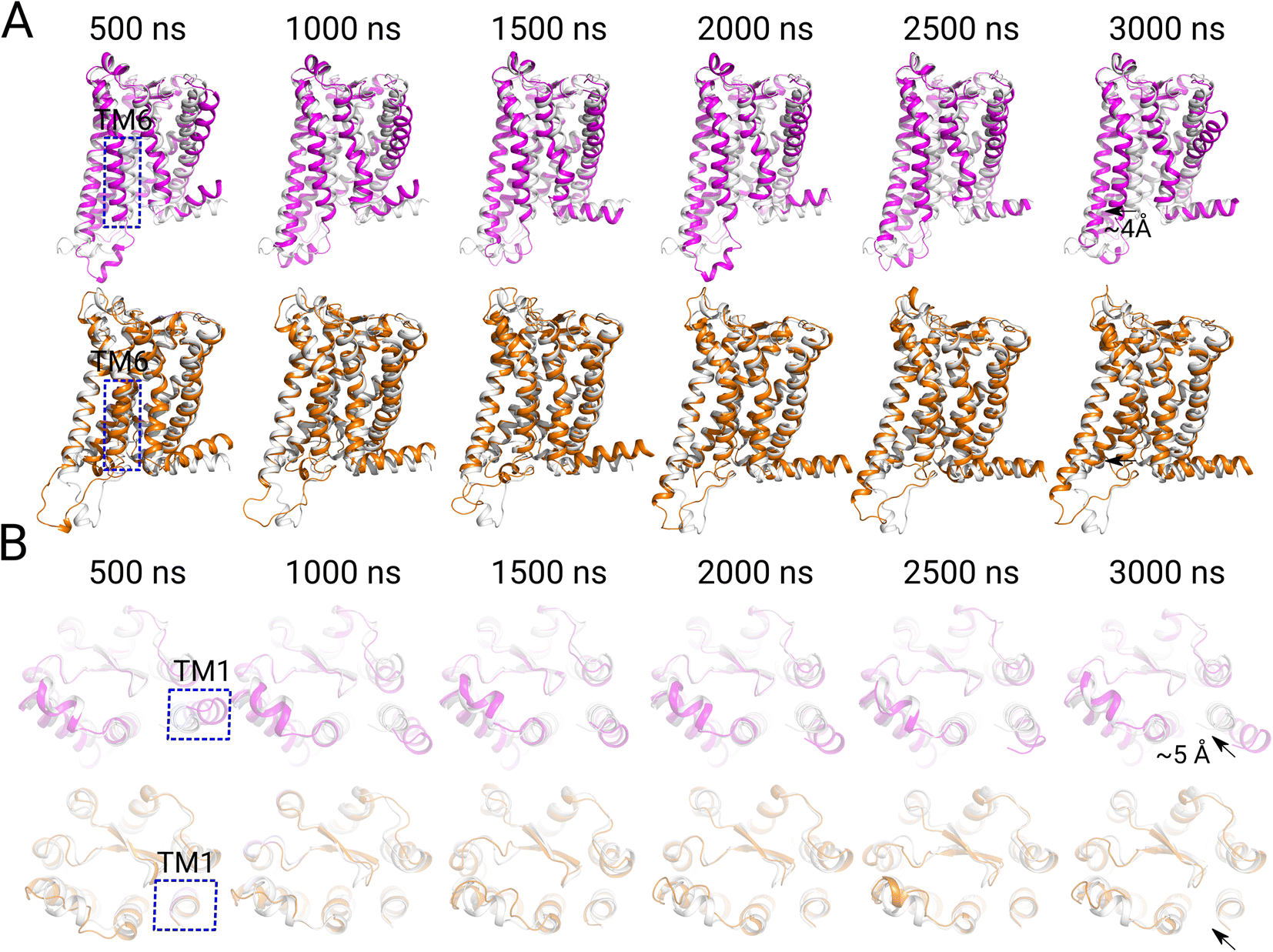
Molecular Basis of Ligand Selectivity for Melatonin Receptors
Wenqiang Cui, Junlin Dong, Shiyu Wang, Horst Vogel, Rongfeng Zou, Shuguang Yuan
- A mechanistic study on ligand selectivity for melatonin receptors.
Full Publication List
2026
- [41] Wenqiang Cui, et al. Decoding xxx catalytic features in enzymes through generative AI. Submitted.
- [40] Wenqiang Cui, et al. De novo design stable and active xxx protein via generative AI. Submitted.
- [39] Wenqiang Cui, et al. Deciphering the molecular basis of magnesium-enhanced xxx receptor activation. Submitted.
- [38] Wenqiang Cui, et al. Synergistic effects of xxx ions with the N-terminus of xxx receptor. Submitted.
2025
- [37] Fanbin Zeng, Cheng Chen, Zhanwei Fu, Haihui Huang, Wenqiang Cui, Yuanyuan Zhou, Yanjie Kong, Xia Liu, Zhiru Xu, et al., A novel beta-carboline alkaloid derivative targeting MDM2-p53 pathway suppresses colorectal cancer progression, Cellular Oncology, 2025, 48(6), 1821-1836.
- [36] Yuzheng Wu, Xiaoxue Ren, Dezhi Xiao, Pei Liu, You Ke, Wenqiang Cui#, Paul K. Chu#, Huaiyu Wang#, Sacrificing Knight for Pawn-Queen Promotion: Sonosensitive Nano-Xanthiums Orchestrate NO to Retain H2O2 for Antibiofilm Therapy, Advanced Materials, 2025, e21809.
- [35] R. Chitsazi, Y. Wu, GPCR Dock 2021 participants, R. C. Stevens, S. Zhao, et al., The 4th GPCR Dock: assessment of blind predictions for GPCR-ligand complexes in the era of AlphaFold, bioRxiv, 2025, 2025.04.18.647407.
- [34] Qianwei Qu, Zhan Zhu, Meiying Zhao, Haifeng Wang, Wenqiang Cui, Xingyu Huang, Zhigang Yuan, Yuting Zheng, Nanyang Dong, et al., Optimization ultrasonic-assisted aqueous two-phase extraction of glabridin from licorice root and its activity against the foodborne pathogen MRSA, Food Chemistry: X, 2025, 26, 102338.
- [33] Qianwei Qu, Xingyu Huang, Zhan Zhu, Jing Wang, Meiying Zhao, Wenqiang Cui, Yuting Zheng, Yaqing Liu, Xueying Chen, et al., Targeting membrane integrity and imidazoleglycerol-phosphate dehydratase: Sanguinarine multifaceted approach against Staphylococcus aureus biofilms, Phytomedicine, 2025, 138, 156428.
- [32] Natalia Gelfand1, Vojtech Orel1, Wenqiang Cui1, Jiri Damborsky, Chenglong Li, Zbynek Prokop, Wen Jun Xie, Arieh Warshel, Biochemical and Computational Characterization of Haloalkane Dehalogenase Variants Designed by Generative AI: Accelerating the SN2 Step, Journal of the American Chemical Society, 2025, 147(3), 2747-2755.
2024
- [31] Wenqiang Cui, Shuguang Yuan, Will the hype of automated drug discovery finally be realized?, Expert Opinion on Drug Discovery, 2024, 19(3), 259-262.
- [30] Chao Liu1, Wenqiang Cui1, Kongfu Zhu, Shuguang Yuan, Liang Sun, Yujie Liang, Jianping Lu, Da Li, Zhiqin Deng, Li Duan, Weiming Zhang, Xiaohai Yu, Daping Wang, Huawei Zhang, Inhibitor screening for volume-sensitive LRRC8A chloride channel, Journal of Biomolecular Structure and Dynamics, 2024, 42(23), 12993-13001.
- [29] Junlin Dong, Shuai Wang, Wenqiang Cui, Xiaolin Sun, Hao Guo, Haotian Yan, Horst Vogel, Zhe Wang, Shuguang Yuan, Machine learning deciphered molecular Mechanistics with accurate kinetic and thermodynamic prediction, Journal of Chemical Theory and Computation, 2024, 20(11), 4499-4513.
- [28] Beatrice Introini1, Wenqiang Cui1, Xiaowei Chu1, Yanyan Zhang, Ana Catarina Alves, Lisa Eckhardt-Strelau, Sebastian Golusik, et al., Structure of tetrameric forms of the serotonin-gated 5-HT3A receptor ion channel, The EMBO Journal, 2024, 43(20), 4451.
- [27] Beimi Cui, Qiaona Pan, Wenqiang Cui, Ying Wang, V. I. P. Loake, Shuguang Yuan, Feng Liu, Gary J. Loake, S-nitrosylation of a receptor-like cytoplasmic kinase regulates plant immunity, Science Advances, 2024, 10(11), eadk3126.
- [26] Haijian Li1, Xiaolin Sun1, Wenqiang Cui1, Marc Xu, Junlin Dong, Babatunde Edukpe Ekundayo, Dongchun Ni, Zhili Rao, Liwei Guo, Henning Stahlberg, Shuguang Yuan, Horst Vogel, Computational drug development for membrane protein targets, Nature Biotechnology, 2024, 42(2), 229-242.
- [25] Chenyang Wu, Marc Xu, Junlin Dong, Wenqiang Cui, Shuguang Yuan, The structure and function of olfactory receptors, Trends in Pharmacological Sciences, 2024, 45(3), 268-280.
2023
- [24] Wenqiang Cui, Junlin Dong, Shiyu Wang, Horst Vogel, Rongfeng Zou, Shuguang Yuan, Molecular basis of ligand selectivity for melatonin receptors, RSC Advances, 2023, 13(7), 4422-4430.
- [23] Tingting Ma, Lin Wang, Anan Chai, Chunhui Liu, Wenqiang Cui, Shuguang Yuan, S. Wing Ngor Au, Lin Sun, et al., Cryo-EM structures of ClC-2 chloride channel reveal the blocking mechanism of its specific inhibitor AK-42, Nature Communications, 2023, 14(1), 3424.
- [22] Qianwei Qu1, Wenqiang Cui1, Xingyu Huang1, Zhan Zhu, Yixian Dong, Zhigang Yuan, Chunhui Dong, Yuting Zheng, Xueying Chen, et al., Gallic Acid Restores the Sulfonamide Sensitivity of Multidrug-Resistant Streptococcus suis via Polypharmaceology Mechanism, Journal of Agricultural and Food Chemistry, 2023, 71(18), 6894-6907.
2022
- [21] Mo Chen, Yan Li, Shuang Li, Wenqiang Cui, Yonghui Zhou, Qianwei Qu, Ruixue Che, Lijun Li, Shuguang Yuan, Xin Liu, Molecular Mechanism of Staphylococcus xylosus Resistance Against Tylosin and Florfenicol, Infection and Drug Resistance, 2022, 15, 6165-6176.
- [20] Lu Zhu, Shiyu Lin, Wenqiang Cui, Yajuan Xu, Lingling Wang, Zhe Wang, Shuguang Yuan, Yiyun Zhang, Ying Fan, et al., A nanomedicine enables synergistic chemo/photodynamic therapy for pancreatic cancer treatment, Biomaterials Science, 2022, 10(13), 3624-3636.
- [19] Shiyu Wang, Xiaolin Sun, Wenqiang Cui, Shuguang Yuan, MM/PB(GB)SA benchmarks on soluble proteins and membrane proteins, Frontiers in Pharmacology, 2022, 13, 1018351.
- [18] Chen Xing1, Wenqiang Cui1, Yue Zhang, Xin-Shu Zou, Jing-You Hao, Sheng-Di Zheng, Ting-Ting Wang, et al., Ultrasound-assisted deep eutectic solvents extraction of glabridin and isoliquiritigenin from Glycyrrhiza glabra: Optimization, extraction mechanism and in vitro bioactivities, Ultrasonics Sonochemistry, 2022, 83, 105946.
2021
- [17] Wenqiang Cui, Fei Yu, Yiyun Zhang, Xue Han, Rongfeng Zou, Yang Tang, Lingling Wang, Nsabimana Eliphaz, Jing Wang, et al., Discovery of traditional Chinese medicines against porcine reproductive and respiratory syndrome virus, Pharmacological Research - Modern Chinese Medicine, 2021, 1, 100003.
- [16] Chun-Liu Dong, Yue Qin, Jin-Xin Ma, Wenqiang Cui, Xing-Ru Chen, Li-Ya Hou, Xue-Ying Chen, Bello-Onaghise God’spower, Nsabimana Eliphaz, Jun-Jie Qin, Wen-Xin Guo, Wen-Ya Ding, Yanhua Li, The Active Ingredients Identification and Antidiarrheal Mechanism Analysis of Plantago asiatica L. Superfine Powder, Frontiers in Pharmacology, 2021, 11, 612478.
- [15] Qianwei Qu1, Wenqiang Cui1, Xiaoxu Xing1, Rongfeng Zou, Xingyu Huang, Xiaoting Wang, Tao Wu, Bello-Onaghise God’spower, et al., Rutin, A Natural Inhibitor of IGPD Protein, Partially Inhibits Biofilm Formation in Staphylococcus xylosus ATCC700404 in vitro and in vivo, Frontiers in Pharmacology, 2021, 12, 728354.
2020
- [14] Jing Wang, Qianwei Qu, Xin Liu, Wenqiang Cui, Fei Yu, Xueying Chen, Xiaoxu Xing, Yonghui Zhou, Yu Yang, et al., 1-Hydroxyanthraquinone exhibited antibacterial activity by regulating glutamine synthetase of Staphylococcus xylosus as a virulence factor, Biomedicine & Pharmacotherapy, 2020, 123, 109779.
- [13] Chun-Liu Dong, Sheng-Di Zheng, Yan-Yan Liu, Wenqiang Cui, Mei-Qi Hao, Bello-Onaghise God’spower, Xue-Ying Chen, et al., Albendazole solid dispersions prepared using PEG6000 and Poloxamer188: formulation, characterization and in vivo evaluation, Pharmaceutical Development and Technology, 2020, 25(9), 1043-1052.
- [12] Ke Liu1, Rongfeng Zou1, Wenqiang Cui1, Meiqing Li, Xueying Wang, Junlin Dong, Hongchun Li, Hongpei Li, Peihui Wang, Ximing Shao, et al., Clinical HDAC inhibitors are effective drugs to prevent the entry of SARS-CoV2, ACS Pharmacology & Translational Science, 2020, 3(6), 1361-1370.
- [11] Bello-Onaghise God’spower, Guangyi Wang, Xue Han, Eliphaz Nsabimana, Wenqiang Cui, Fei Yu, Yiyun Zhang, et al., Antiviral strategies of Chinese herbal medicine against PRRSV infection, Frontiers in Microbiology, 2020, 11, 1756.
- [10] Wenqiang Cui, Adnane Aouidate, Shouguo Wang, Qiuliyang Yu, Yanhua Li, Shuguang Yuan, Discovering Anti-Cancer Drugs via Computational Methods, Frontiers in Pharmacology, 2020, 11, 733.
2019
- [9] Jing-Wen Bai, Xing-Ru Chen, Yang Tang, Wenqiang Cui, Dong-Li Li, Bello-Onaghise God’spower, Yu Yang, Study on microwave assisted extraction of chrysophanol and its intervention in biofilm formation of Streptococcus suis, RSC Advances, 2019, 9(50), 28996-29004.
- [8] Wenqiang Cui, Qian-Wei Qu, Jin-Peng Wang, Jing-Wen Bai, Bello-Onaghise God’spower, Yan-An Li, Yong-Hui Zhou, et al., Discovery of Potential Anti-infective Therapy Targeting Glutamine Synthetase in Staphylococcus xylosus, Frontiers in Chemistry, 2019, 7, 381.
- [7] Qianwei Qu, Jing Wang, Wenqiang Cui, Yonghui Zhou, Xiaoxu Xing, Ruixue Che, Xin Liu, Xueying Chen, et al., In vitro activity and In vivo efficacy of Isoliquiritigenin against Staphylococcus xylosus ATCC 700404 by IGPD target, PLoS One, 2019, 14(12), e0226260.
- [6] Xin Liu, Jing Wang, Mo Chen, Ruixue Che, Wenya Ding, Fei Yu, Yonghui Zhou, Wenqiang Cui, Xiaoxu Xing, et al., Comparative proteomic analysis reveals drug resistance of Staphylococcus xylosus ATCC700404 under tylosin stress, BMC Veterinary Research, 2019, 15(1), 224.
- [5] Yan-Yan Liu, Xing-Ru Chen, Jin-Peng Wang, Wenqiang Cui, Xiao-Xu Xing, Xue-Ying Chen, Wen-Ya Ding, et al., Transcriptomic analysis reveals flavonoid biosynthesis of Syringa oblata Lindl. in response to different light intensity, BMC Plant Biology, 2019, 19(1), 487.
- [4] Wen-Ya Ding, Sheng-Di Zheng, Yue Qin, Fei Yu, Jing-Wen Bai, Wenqiang Cui, Tong Yu, Xing-Ru Chen, et al., Chitosan grafted with beta-cyclodextrin: synthesis, characterization, antimicrobial activity, and role as absorbefacient and solubilizer, Frontiers in Chemistry, 2019, 6, 657.
2018
- [3] Yan-Yan Liu1, Xing-Ru Chen1, Ling-Fei Gao, Mo Chen, Wenqiang Cui, Wen-Ya Ding, Xue-Ying Chen, Bello-Onaghise God’spower, Yanhua Li, Spectrum-Effect Relationships Between the Bioactive Ingredient of Syringa oblata Lindl. Leaves and Its Role in Inhibiting the Biofilm Formation of Streptococcus suis, Frontiers in Pharmacology, 2018, 9, 570.
- [2] Wenya Ding, Yonghui Zhou, Qianwei Qu, Wenqiang Cui, Bello-Onaghise God’spower, Yan-Yan Liu, Xing-Ru Chen, Mo Chen, et al., Azithromycin Inhibits Biofilm Formation by Staphylococcus xylosus and Affects Histidine Biosynthesis Pathway, Frontiers in Pharmacology, 2018, 9, 740.
2017
- [1] Xing-Ru Chen, Xiao-Ting Wang, Mei-Qi Hao, Yong-Hui Zhou, Wenqiang Cui, Xiao-Xu Xing, Chang-Geng Xu, Jing-Wen Bai, Yanhua Li, Homology Modeling and Virtual Screening to Discover Potent Inhibitors Targeting the Imidazole Glycerophosphate Dehydratase Protein in Staphylococcus xylosus, Frontiers in Chemistry, 2017, 5, 98.
📘 Patents
- [8] Li Yanhua, Ding Wenya, Qin Yue, Wenqiang Cui, Yan Qingbo, Hou Liya, Liu Yanyan, Dong Chunliu. Anti-diarrhea active ingredients of plantain and their application. Application No.: CN201911114532.7. Grant No.: CN111170977B. Authorized on 2024.11.08.
- [7] Bai Jingwen, Tang Yang, Yang Yu, Li Yanhua, Bai Xuedong, Wenqiang Cui, Xu Yaqin. Use of honokiol in inhibiting Streptococcus suis or its biofilm. Application No.: CN202010137016.2. Grant No.: CN111419829B. Authorized on 2023.08.18.
- [6] Shuguang Yuan, Xianchong Chen, Mupeng Luo, Wenqiang Cui, Rongfeng Zou, Jincun Zhao, Jing Sun. Application of bacillamide A in the preparation of drugs for the prevention and treatment of coronavirus infection. Application No.: CN202010265260.7. Grant No.: CN111420024B. Authorized on 2023.10.03.
- [5] Shuguang Yuan, Wenqiang Cui, Mupeng Luo, Rongfeng Zou, Xianchong Chen, Jincun Zhao, Jing Sun. Application of pyroglutamic acid in the preparation of drugs for the prevention and treatment of novel coronavirus infection. Application No.: CN202010266183.7. Grant No.: CN111467338B. Authorized on 2021.04.23.
- [4] Shuguang Yuan, Wenqiang Cui, Mupeng Luo, Rongfeng Zou, Xianchong Chen, Jincun Zhao, Jing Sun. Application of sodium phosphonoformate in the preparation of drugs for the prevention and treatment of coronavirus infection. Application No.: CN202010265267.9. Grant No.: CN111467355B. Authorized on 2021.04.23.
- [3] Yanhua Li, Wenqiang Cui, Yonghui Zhou, Qianwei Qu, Jinping Wang. Use of sorafenib in antibacterial applications and intervention against pathogenic bacterial biofilms. Application No.: 201811047450.0. Grant No.: CN108785306B. Authorized on 2021.02.19.
- [2] Yanhua Li, Jinping Wang, Wenqiang Cui, Qianwei Qu, Xin Liu, Fei Yu, Yonghui Zhou, Xiaoxu Xing. Use of 1-hydroxyanthraquinone in the preparation of antibacterial agents, agents for interfering with pathogenic bacterial biofilms, and drugs for bovine mastitis. Application No.: ZL201110263589.0. Grant No.: CN108969510B. Authorized on 2021.02.19.
- [1] Yanhua Li, Qianwei Qu, Wenqiang Cui, Jinping Wang, Yonghui Zhou, Ruixiang Che, Xiaoxu Xing. Use of isoliquiritigenin in antibacterial applications, biofilm intervention, and treatment of bovine mastitis. Application No.: CN201811047444.5. Grant No.: CN109200038B. Authorized on 2021.02.12.
🏅 Honors and Awards
- Reviewer for Nature Communications, Journal of Agricultural and Food Chemistry, Frontiers in Pharmacology, Frontiers in Chemistry, and Frontiers in Microbiology, Computational and Structural Biotechnology Journal.
- Reviewer Editor for Frontiers in Pharmacology (Experimental Pharmacology and Drug Discovery).
- Guest Editor for Membranes.
- Author of highly cited articles in computational drug discovery and molecular design.
- Member of the Chinese Chemical Society.
- Member of Sigma Xi, the U.S. honor society for scientific research.
📬 Contact
Email: wenqiang.cui “AT” ufl.edu